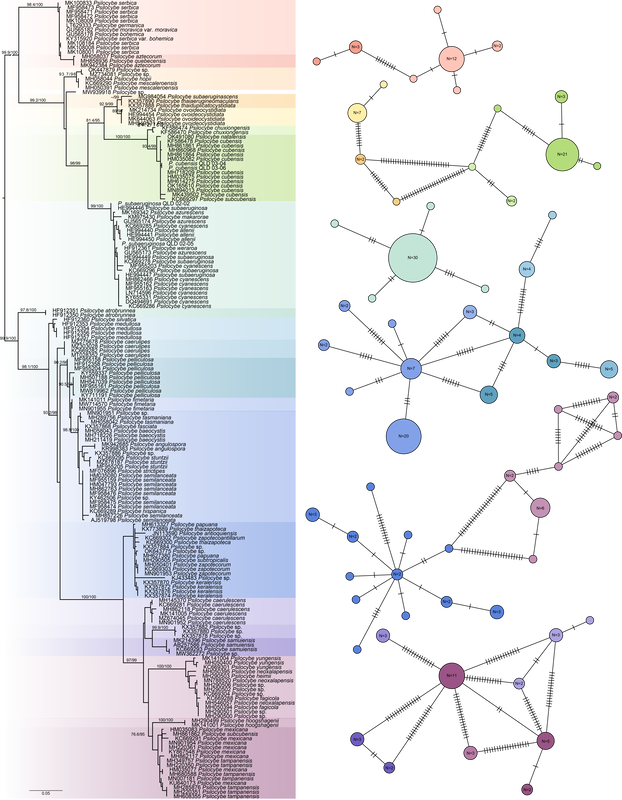
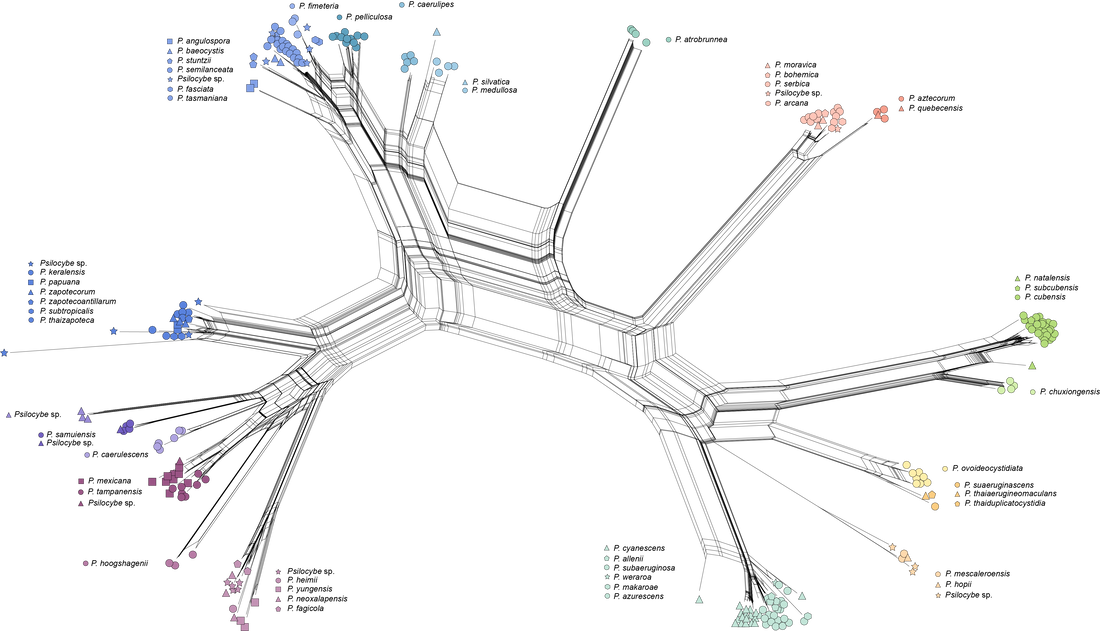
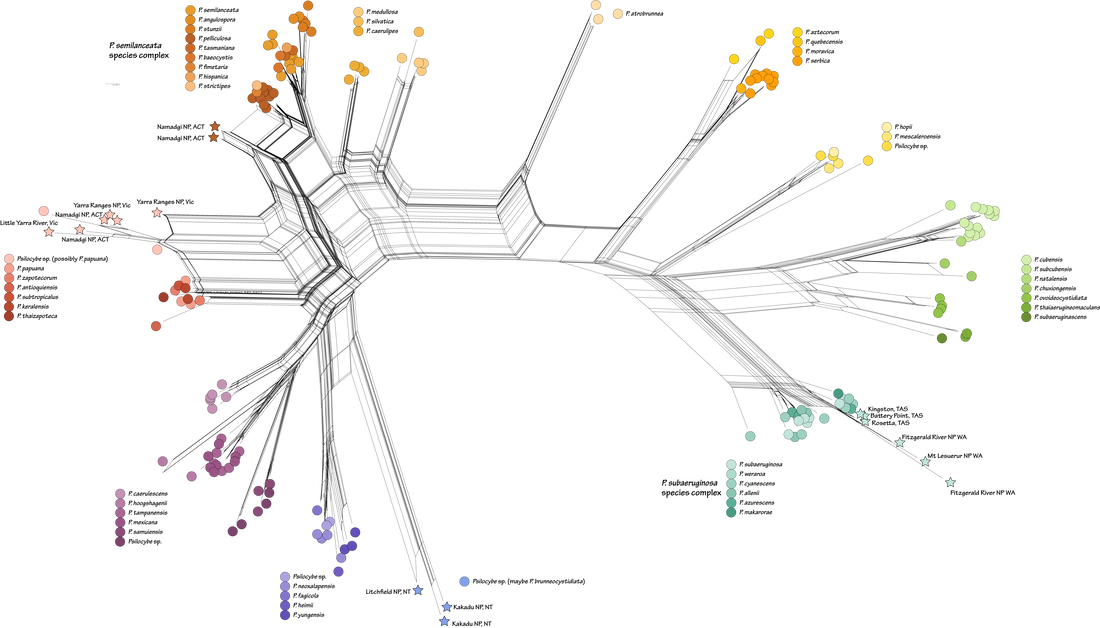Sequences from metabarcoding data: A 'new' hallucinogenic species occurs in the Northern Territory1/18/2022 Yesterday I guest lectured in Liz Aitken's Fungal Biology class at UQ (BIOL3210). Fun on a bun. One student asked a question along the lines of 'are gold tops only present in cow farms?' I answered that they are potentially everywhere but need certain conditions to fruit, this is based on their ability to distribute spores and that saprotrophic fungi could form hyphae wherever there is a source of carbohydrates they can access. How to test this question a bit more? I jumped on to Bioplatforms Australia and downloaded the curated fungal ITS database, which includes all ITS sequences (~250 base pairs) from sampling sites around Australia (link to the data here). I expected that P. cubensis would occur in many sampled environments, even if there were no cows around. How wrong I was. There are 3,412 BASE sample sites. Many of the fungi identified as Psilocybe in their dataset were Deconica, and when these are removed, species of Psilocybe occur at 17 of those sites (<0.5% of sampled sites!). Here's a link to a pdf of the above figure if you want more resolution.
One of the most exciting things to me is the putative hallucinogenic species of Psilocybe in the Northern Territory, recorded in Kakadu and Litchfield National Parks. These sequences, which had just under 100 repeats in the dataset, formed a monophyletic group sister to the Psilocybe yungensis clade, and not closely related to anything else. Potentially a candidate for a new taxon (if species of Psilocybe have not been described from the NT), and potentially evidence that hallucinogenic species of mushroom occur in Kakadu NP. To all those reading in the NT, why not head to Litchfield NP for the day next time you've had some rain :). Send me some spores and we can describe it if it's a new taxon. I've added the location and number of reads to the BASE sequence data. You'll notice there are no sequences of P. cubensis or P. semilanceata in the BASE data, which I find surprising for two 'weedy' taxa in Australia. Plenty of P. subaeruginosa s. lat., with some of the those sequences near-identical to P. weraroa (which is not very informative in this complex). More sequences and locations than I had expected for the P. papuana clade. What does this mean? Psilocybe are present, whether as spores or mycelium, at the sample sites. The sampling method of BASE may have biased whether Psilocybe was sequenced, which may explain the relatively few sites that had Psilocybe present. And lastly, in answer to the question from mycology yesterday, I'll change my answer to maybe P. cubensis is less ubiquitous than I thought :)
0 Comments
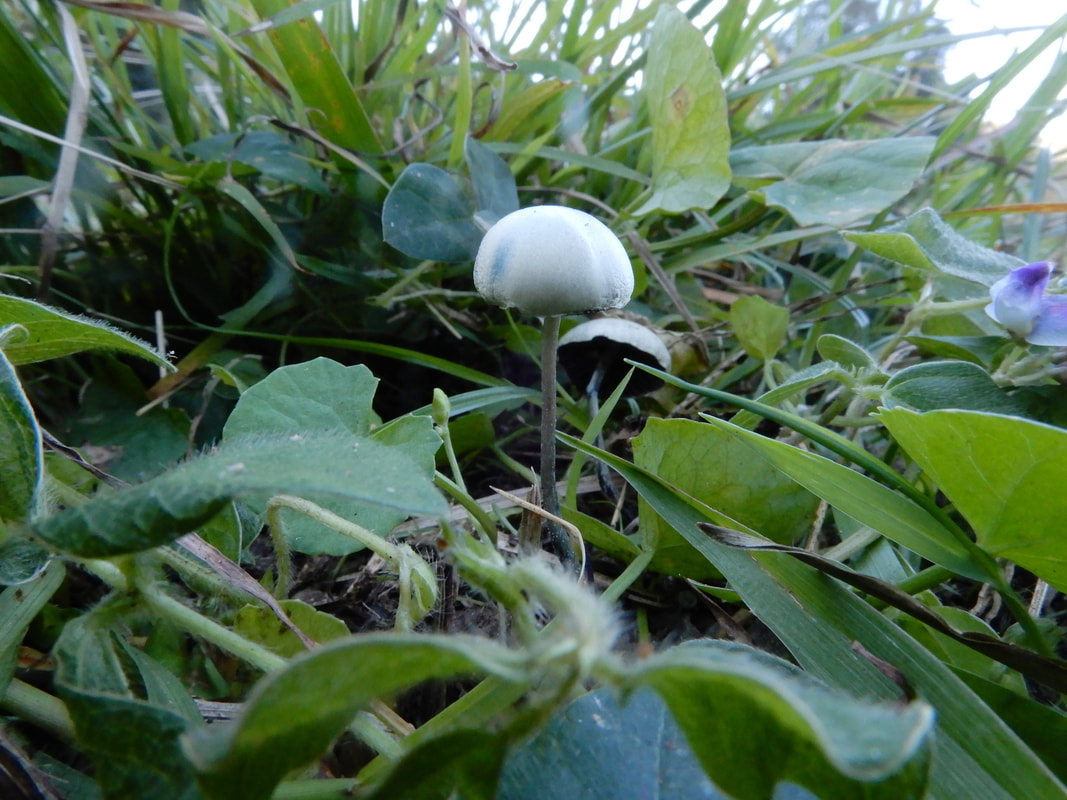
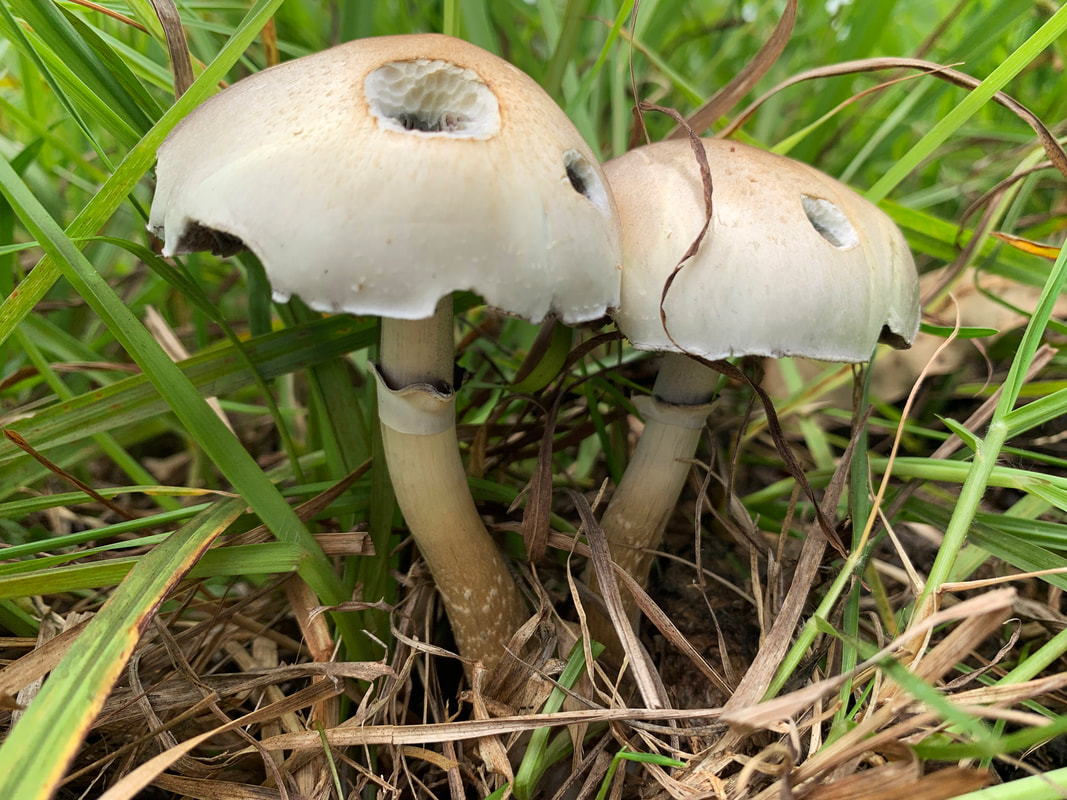
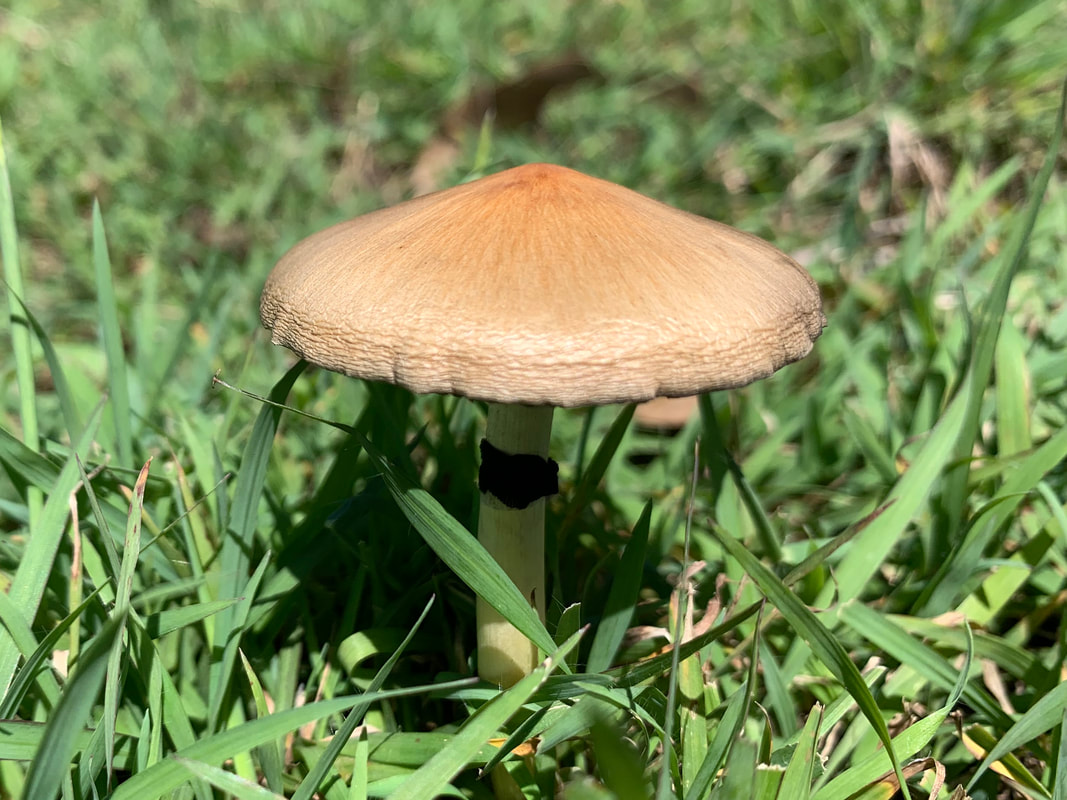
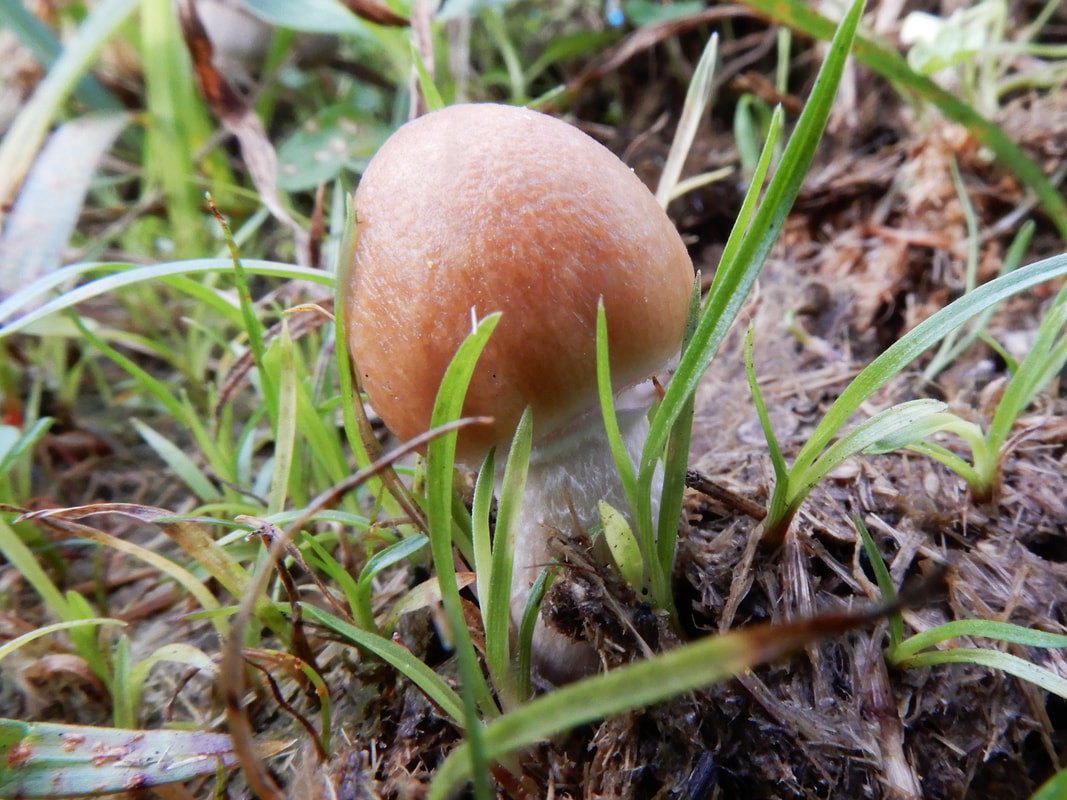
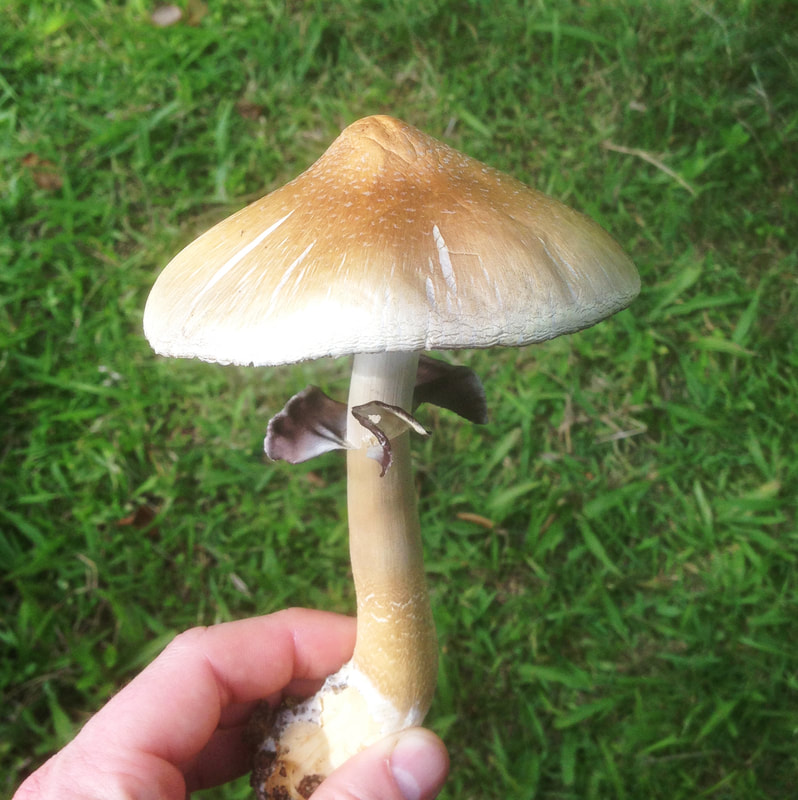
I photographed these Psilocybe cubensis to illustrate what I think is herbivory (and I'm always prepared to be wrong). A chemist colleague of mine was chatting about the origin of psilocybin to deter insects, but I think that hypothesis requires testing.
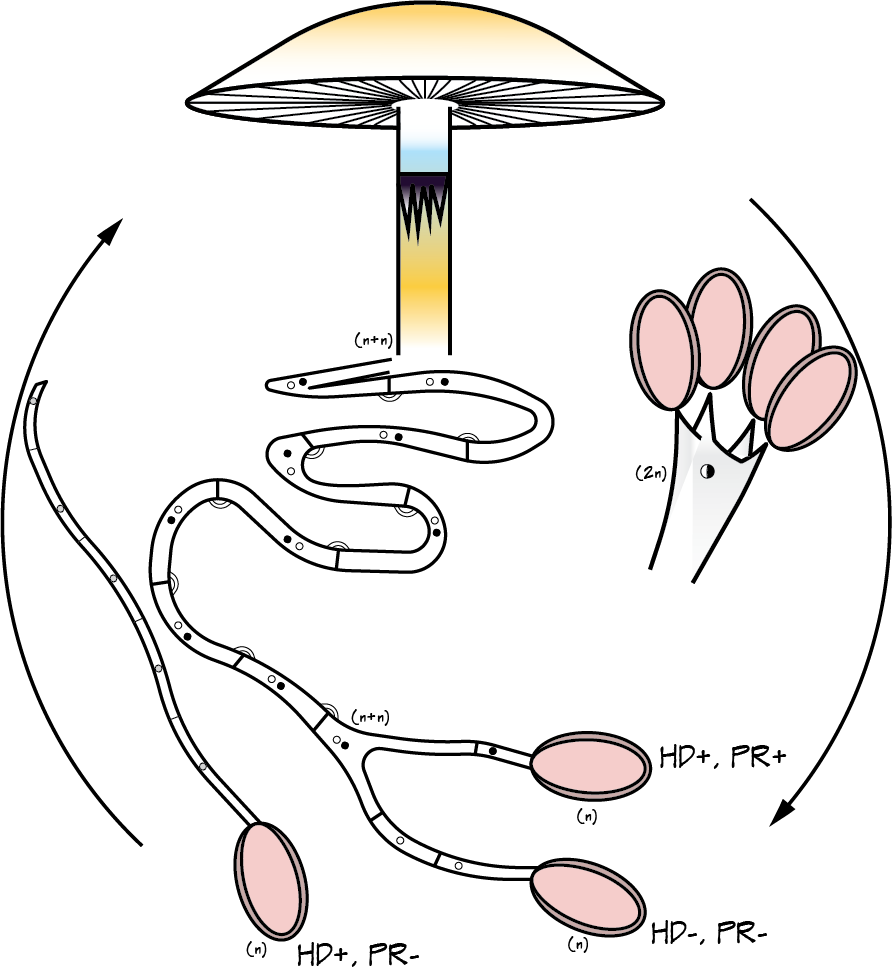
Maybe knowledge of who eats Psilocybe now can shed light on why psilocybin evolved. Whether it was a deterrent or an attractant to help spread spores is something I would give anything to know :). A complex gene pathway that has evolved convergently and been acquired horizontally indicates there is an evolutionary advantage to produce psilocybin. I've also wondered whether cows mistakenly hoover up stray mushrooms from long grass. I can imagine they would, and a farmer thought his cows went 'loopy' every so often. Surely humans aren't the only beneficiaries of psilocybin. All this talk about haploid stages, I thought I should illustrate it for y'all. Mushrooms (n+n) are the site of recombination, which occurs after nuclei fuse in the basidium (karyogamy to make a diploid nucleus, 2n). Basidiospores are haploid (n), containing one recombinant nucleus.
I use a single basidiospore to grow cultures to make the genetic analyses easier with only one copy of each chromosome instead of two in a dikaryon (n+n) or diploid (2n). I've added mating genes in Basidiomycota to the figure; the homeodomain locus (HD), which usually has two genes, and the pheromone/receptor locus. Alleles of all genes at these loci must differ for a compatible cross. I'll cover this more when I've annotated the mating genes in our genomes of P. cubensis and P. subaeruginosa. Thank you to AGRF for the high quality genomic data that we have since assembled. Each sample had 19–27 million reads (about 6–8 Gb of data). I was worried that I didn't sequence Psilocybe, however, our rDNA contigs are a perfect match for P. subaeruginosa and P. cubensis (pictured above). Phew. (The resolution of the tree is atrocious, if you're desperate to see the taxon selection, there is a pdf here). My next worry was that I had contaminated haploid cultures with dikaryotic cultures. What a relief, everything that is meant to be haploid, was haploid. The below figure shows that the one dikaryon (essentially a diploid) we sequenced was a much worse assembly than our haploids. I'll use this information to only sequence haploids in the future. Next step is to improve these assemblies with some long reads from PacBio. Stay tuned for updates on what we pull out from these genomes in the next few weeks... with a decent annotation, we may already be able to make a case that P. cyanescens is con-specific with P. subaeruginosa, depending on their similarity. 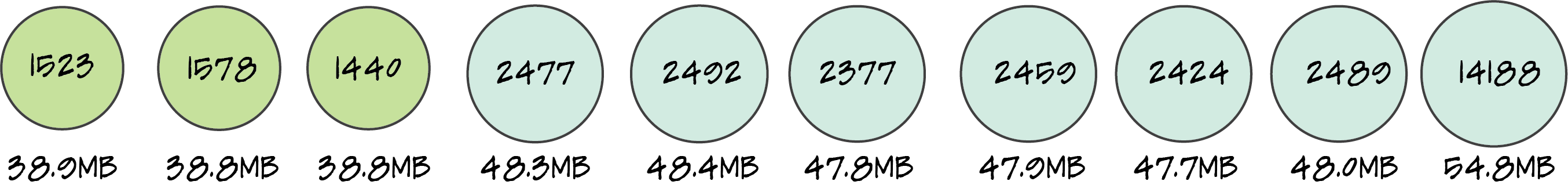 Genome assemblies for Psilocybe cubensis (first three on the left) and P. subaeruginosa. The circle size is proportional to genome size, and number of contigs is centred in circles. The far right assembly was a dikaryotic culture of P. subaeruginosa, all other assemblies were haploid. We want the number in the circle to be as small as possible to reflect the number of chromosomes. Hopefully every potential reader knows I could never ask you to collect mushrooms on my behalf. The penalties for citizen scientists are severe for possessing psilocybin.
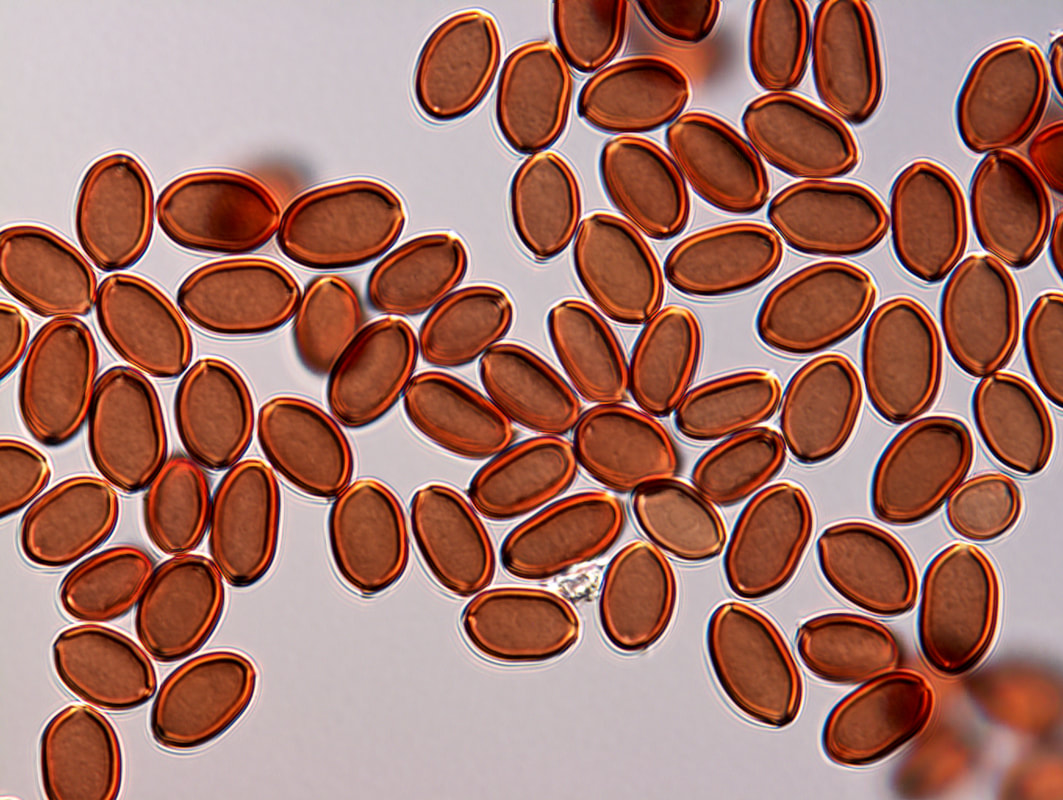
Thank you to those who have sent me spore prints from their own incidental private research or while collecting for herbaria. These high-res images of basidiospores of Psilocybe cubensis are for your personal use (available here). The University of Queensland covers my salary four days a week as a research fellow. We have applied to Advance Queensland, the Australian Biological Resources Study and the Hermon Slade Foundation, but our applications have not been persuasive to fund any research on magic mushrooms. Specimen collection is a huge bottleneck for the project because I don't have resources to travel, and I really appreciate spore contributions to help increase our potential sampling. The data I have acquired and resources I use are funded creatively and on the cheap through my other research projects in plant pathology. For those enthusiastic to provide material, any culture made is lodged in the Queensland Plant Pathology Herbarium with the collector's name (if they are willing). My vision is that if a culture from a citizen scientist is used commercially, there will be a benefit sharing agreement with either the collector or land holder. I won't be the one to commercialise mushrooms, and who knows how far down the track this would be... but you never know. (Interestingly, not a single one of the >24,000 cultures in the Herbarium has been commercialised, maybe these will be a first). Thank you to everyone who has sent me photographs of Psilocybe cubensis fruiting after these November rains. It's fantastic to see the excitement, and hopefully we can answer whether this mushroom is native, or if introduced (which we suspect), how many times it has escaped. Round two submitting samples to AGRF. First attempt directly from agar and stored in >95% ethanol for a month or two did not yield DNA. This time I've grown cultures in liquid (PD broth) for two weeks and submitted to AGRF. We should have some feedback for this approach next week... I'm already optimistic. We've submitted seven haploid cultures of P. subaeruginosa and three of P. cubensis. Plenty of P. cubensis around southeast Queensland at the moment. We're starting to exhaust our sampling locally and it will be time to move further afield. The aim of this part of the project is to determine whether gold tops are introduced, and if yes, how many times. We suspect it is an introduction and will sequence about 50 genomes from 10 or more locations (separated by ≥ 100 kms). If we observe high genetic diversity, we will need to re-think our sampling.
We are about to sequence the genomes of 6 haploid cultures collected from a population of Psilocybe subaeruginosa in Queensland. It will be nice to have some data in hand and we can compare our genomes to those available for P. cyanescens to test whether these taxa are con-specific (the same species).
The haploid cultures have some interesting phenotypes (fast smooth growth, slow and patchy growth, fast radial growth). Knowledge of these phenotypes may one day help with crossing different haploid cultures to tailor genetics. Expect an update when we get sequence data back from AGRF :) Looking forward to the rains to expand sampling of P. cubensis in the next few months. We have the following aims.
1. To determine which mushrooms are native to Australia We will sample widely from populations of P. subaeruginosa in Australia to answer whether populations are structured by geography or substrate. We hypothesise P. subaeruginosa is native to Australia and will have relatively high genetic diversity across sampled populations. If P. subaeruginosa is endemic to Australia, there may be flow-on impacts for patents and commercialisation under the Queensland Biodiscovery Act (2006). We expect introduced taxa to have less genetic diversity than native species. For example, invasive fungi that are dikaryotic/diploid (have two copies of a genome, one chromosome inherited from each parent), will have a maximum of two alleles at one locus (heterozygous) if there has been one introduction. More diversity means more introductions (or high diversity from one introduction), but we expect it to be much lower when compared to a native taxon. For example, we may expect P. cubensis to have 3–5 alleles at a locus, compared to P. subaeruginosa that may have many more! 2. To determine how many species occur in Australia In studying populations of P. subaeruginosa, we will resolve the biodiversity of this taxon, specifically whether there are multiple species in the complex. 3. To determine if there is variability in the gene cluster that produces psilocybin across the sampled diversity of P. subaeruginosa Different mushrooms have different effects on people, for example, someone who dabbles in shamanism may tell you that the experience on blue meanies is different to the experience on gold tops. Is this variation genetic? If yes, will crossing a collection of P. subaeruginosa from Tasmania with one from Queensland have an impact on the ratio or action of psilocybin? We expect greatest genetic diversity in the centre of origin of a taxon, and maybe the therapeutic world will benefit from advances in the genetics of psilocybin production. 4. An outcome of our study is to start the first living, culture collection of Psilocybe to safeguard against habitat loss and as a platform for future research A spore print will be made from every mushroom used in the study, and 5–6 cultures will be made from basidiospores (which have one copy of a genome (haploid)). We will phenotype these cultures for growth rate and, potentially, ability to produce psilocybin. The genome of every culture will be sequenced and from there we can determine mating type, versatility of psilocybin production and any other phenotype we can link to a gene. The cultures get stored in a Herbarium (the largest culture collection in Australia) and if anyone ever commercialises a fungus from the collection, they have to enter a benefit sharing agreement with the original land owner, collector or Queensland Government (if collected in Queensland). Background on Psilocybe subaeruginosa
Our 'flying saucer' mushroom in Australia, Psilocybe subaeruginosa, was described in 1927 by John Burton Cleland. The specimens studied by Cleland were collected from grass or decaying wood in national parks of South Australia, Victoria and New South Wales (there is a link to JB's original description at this site). Presumably JB didn't know this was a hallucinogenic mushroom at the time, and the properties of psilocybin were not published by Hoffman until 1958. Other (non-Australian) mycologists considered there was more biodiversity and described Psilocybe australiana, P. eucalypta and P. tasmaniana in 1978. Non-Australian mycologists have had tremendous impact on the taxonomy of Psilocybe and the community owes much to the passion of Gaston Guzman and Roy Watling (if you're a Brisbane local I recommend reading Watling's account of gold tops growing in Indooroopilly). The key take home from all that is P. subaeruginosa and related taxa (whether they are more than one species or not) occur in natural areas of Australia, are diverse and were described nearly 100 years ago. A species of Psilocybe, P. cyanescens, has become prolific in the northern hemisphere (Europe and the United States). It grows in wood chips in non-natural areas. It was first described from Kew Botanic Gardens in 1946 (link to the original description here). This mushroom, along with P. cubensis, have become the standards for research on psilocybin and are the basis for many patents on extraction, production and therapeutic use of psilocybin (check out a list of psilocybin patents here). Our hypothesis is that P. subaeruginosa and P. cyanescens are the same species, and that Australia is the likely centre of origin of this taxon. This is based on their years of description (keep in mind that Australian mycology lags behind the northern hemisphere), the habitat of these mushrooms (natural areas of Australia, non-natural areas northern hemisphere) and their near-identical sequences of a barcoding locus of DNA. At some point before 1946, soil or wood chips from Australia may have been transported to Kew or botanic gardens globally, and inadvertently spread P. subaeruginosa, spawning its success as an invasive fungus. Background on Psilocybe cubensis Speaking of prolific mushrooms... gold tops are widespread along the east coast of Australia. They grow out of cow manure as far up as the Daintree (or this is where I've collected them in northern Queensland, they could be even further north). How did they get to Australia and become so successful in this niche? One hypothesis is they are native here, perhaps once dung fungi of Diprotodons that wandered the landscape. This does not seem likely. Gaston Guzman treated P. cubensis as a native of Central America, and he noticed similarities in the morphology of species of Psilocybe in different geographic areas, Our hypothesis is they were introduced to Australia. It will be fascinating to know how many times and whether one introduction is the source of their proliferation along the east coast. Spores of P. cubensis are shared internationally, perhaps more genetic diversity has escaped into Australia from enthusiasts who have released spores. Does all this reasoning make you yearn for answers? Are flying saucer mushrooms endemic to Australia, and just how many species do we have? Are gold tops native to Australia, if not, how many times have they 'escaped'? If mushrooms have been present in Australia longer than humans, is there a chance people have known about their properties here since before 1958 (when Hoffman described psilocybin)? Welcome to my world if you want to know the answers to these questions :) |
Designer Shrooms @ Funky Fungus on 1st July 2023
I started a gig at Funky Fungus as Chief Scientific Officer to make designer shrooms Our research on Psilocybe
|