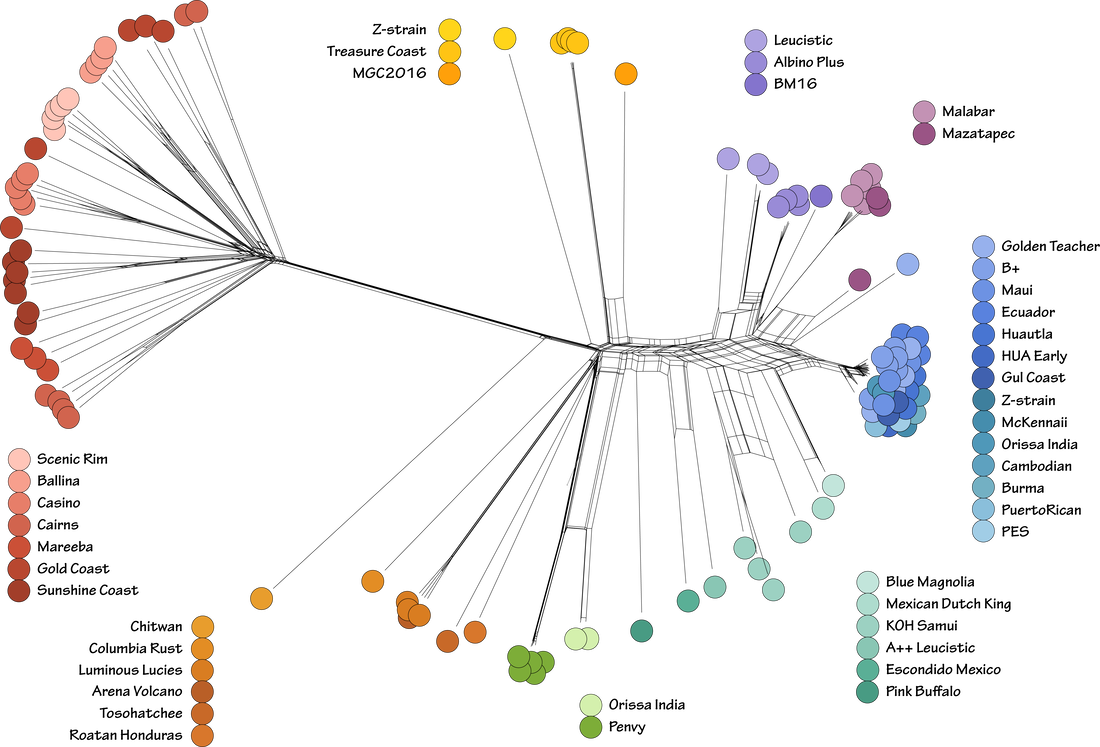|
I collected a couple of Psilocybe cubensis before the season commences because these might have alleles that are suited to fruiting in versatile conditions. Could be interesting to incorporate some of these genetics in future crosses.
The cultures for genomes have grown and I will take samples to AGRF this week. We could expect genome data by Christmas, and with that we'll have a good indication of species diversity in Australia (to answer the question of whether Psilocybe subaeruginosa is more than one taxon). We're submitting a manuscript on mating in P. subaeruginosa, and if that gets through review, I can share some neato knowledge about the psilocybin gene cluster in wood loving species. More answers on cubes coming soon as well.
0 Comments
Thank you to Kelly from BioPlatforms Australia, who has helped support sequencing of 100 genomes of Psilocybe subaeruginosa collected in 2022. Thank you also to the collectors of P. subaeruginosa: Caine B (and helpers), Snu, Jan T, Dave H (and team from Kosciuszko), Tim S, and anyone who has anonymously sent prints. Our project is just over one year in, with quite a lot of data considering there has been no financial support (until BioPlatforms stepped in). There are over 100 haploid cultures of Psilocybe lodged in the herbarium, and we will have sequenced 140 genomes in less than 18 months. This is an effort from the community interested in Psilocybe, I'm just the bloke at the end of the chain.
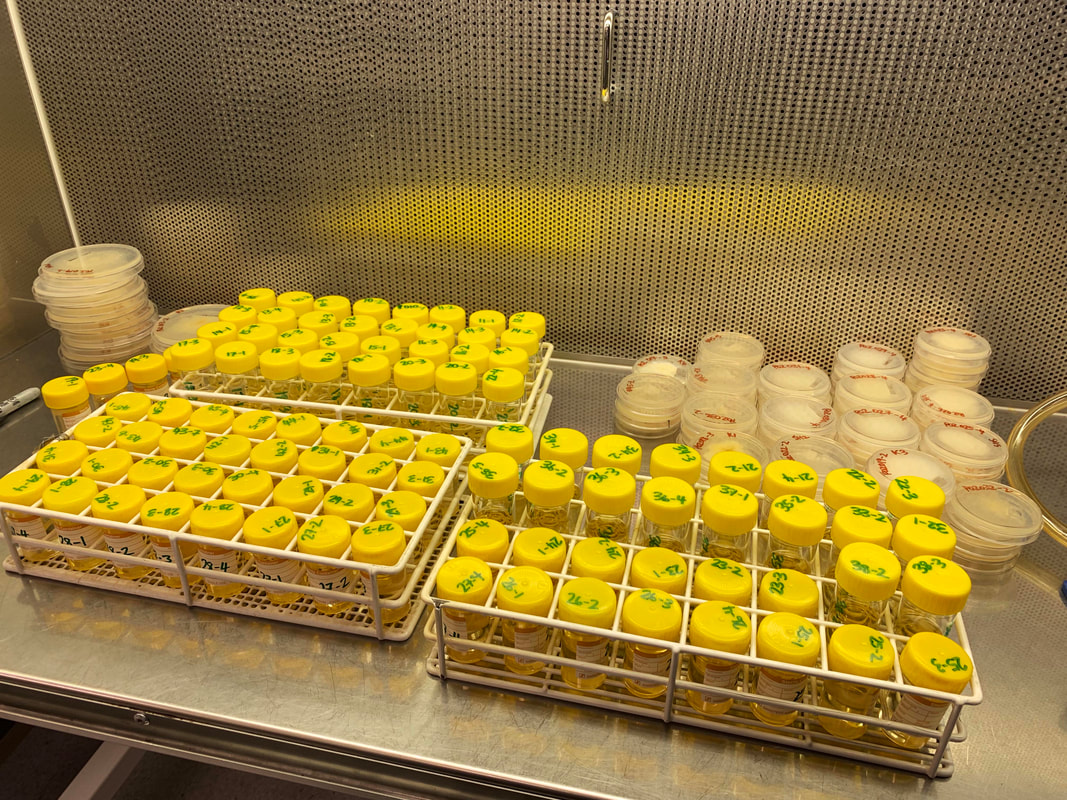
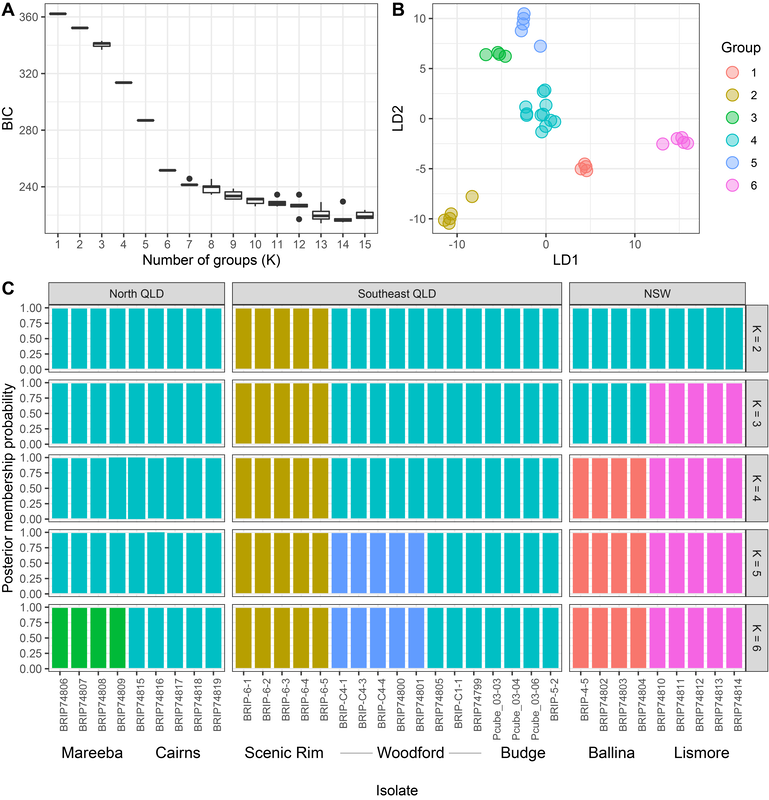
The stories on mating, biodiversity, introduction of P. cubensis, and variation at the psilocybin locus are unfolding nicely. I probably won't have finished everything before my contract finishes in November, but I'll keep working on genetics of magic mushrooms as long as I have a computer, and all these data will be made public for the community to blaze ahead. Pictured below, preparation of the 100 cultures for sequencing. Get in touch if you would like to make use of the data, otherwise they'll be public in 2 years maximum (from when I get the raw data). Here are some new analyses with a longer-term aim to determine how long gold tops have been in Australia. Originally, in the network analyses from past posts, it appeared that populations of gold tops were structured by geography, e.g. mushrooms from Mareeba would not reproduce with mushrooms from Lismore. On the road to determine the effective population size of Psilocybe cubensis in Australia, it now looks like there is some admixture (sexual reproduction among populations). The below figure is a journey to determine how we should treat the populations of P. cubensis in Australia. I tested how many groups best fit the data (which is 'K' in the figure). The figure depicts how populations across Queensland and NSW would look if treated as 2–6 different groups. I settled on 6 as the number of groups in the sequenced populations. If we added new locations, no doubt we would see more geographic structure. The curious result is that populations from Cairns, Gold Coast and Sunshine Coast all group together. They are genetically similar. This could mean that manure has been moved from southeast Queensland to northern Queensland (or vice versa). Though it is difficult to know for sure. It's also interesting that different mushrooms from the Sunshine Coast are either related to those in Cairns, or form their own segregated population.
Leave it with me, but if you have ideas on why these three locations may not be structured by geography, let's explore them together :). Thanks again to the legends who collected the mushrooms for these analyses! Here you can visualise diversity of the psilocybin gene cluster in populations of gold tops in NSW and Queensland. We could pretty much say there are four alleles of the psilocybin gene cluster present across the distribution of gold tops. There is near 100% sequence identity across 30,000 base pairs that encode four genes for psilocybin production. The diversity occurs mostly in two genes, PsiK and PsiM. Whether this has an impact on psilocybin phenotype is something I intend to find out through breeding.
I produced this figure for a talk at Splendour in the Grass coming up on the 23rd of July. Come and say hi if you're going to be there. We potentially have funding to sequence the 120 or so cultures I've made from spore prints sent in from you all in the community. I'll share more when it's official. If you have a population of Psilocybe subaeruginosa that you want included in the study, I'll need those spores in the next couple of weeks. Get in touch if you want to know how your population compares to others in Australia (we have good sampling from Tassie, Victoria, and southeast Queensland). Below is a new way to visualise the diversity of gold tops in Australia compared to the populations cultivated globally. Note that there is not much diversity in the Golden Teacher strain. Our analyses of this show it has a low effective population size (probably from 50+ years of inbreeding) especially compared to the diversity in our outbreeding population here in Australia. More on this hopefully coming in a publication with the Oz genomes soon. Here's a taste of what will come from 32 new Psilocybe cubensis genomes freshly assembled. First of all, thank you to Andrew B, Matt W, Bocky and Dan, Rhys L, and all landholders in southeast Queensland for spores. We ended up with over 50 haploid (or so I thought) cultures of Psilocybe cubensis.
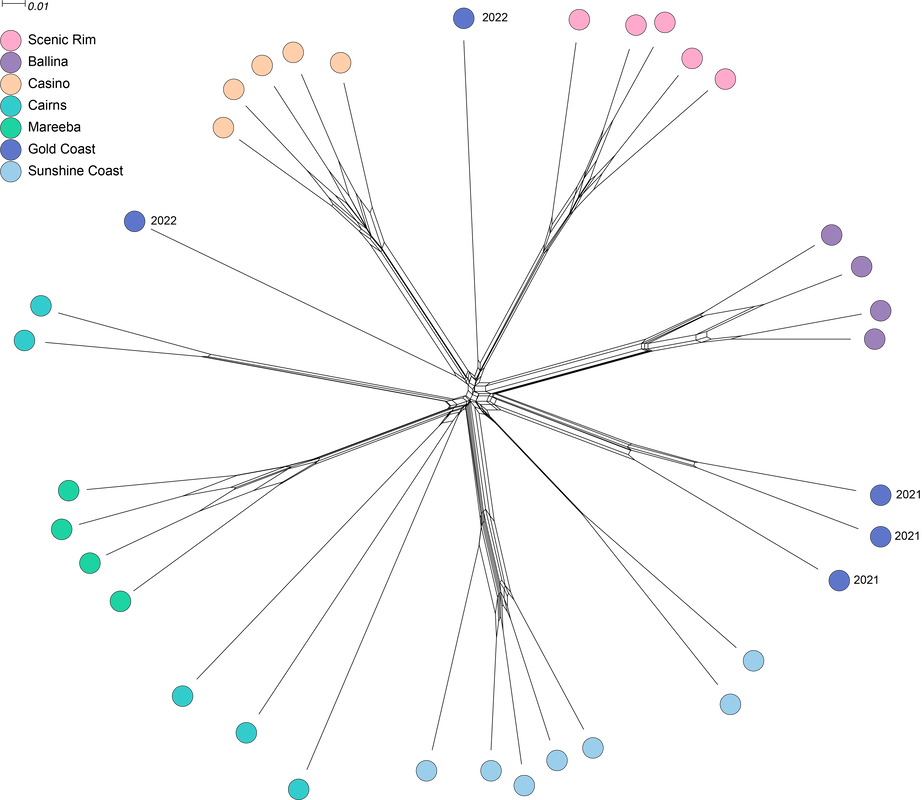
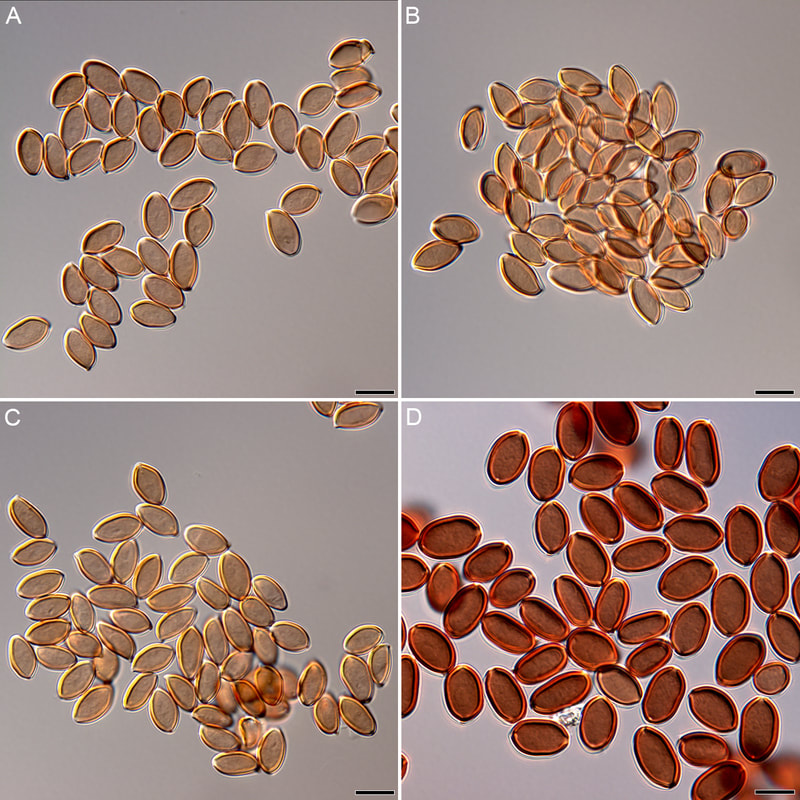
Three of the 32 genomes were probably dikaryotic, which is a reflection on how good I think I am at culturing compared to my actual culturing ability. The figure is a network based on 529,027 SNPs. Preliminary interpretation, there is probably structure by geography, the populations are not admixed. More soon, including some answers about mating! We've resolved reproduction in Psilocybe, now just going through the motions of publishing. This figure compares basidiospores from P. subaeruginosa collected in SE Queensland (A & C) and Tasmania (B, by Caine B), and of P. cubensis (D). Psilocybe cubensis and P. subaeruginosa are sister taxa (they shared a most recent common ancestor). One might interpret there is not much morphological difference between the populations of P. subaeruginosa separated by a strait and two states.
Recording available here: https://vimeo.com/713129405/2c456a2298
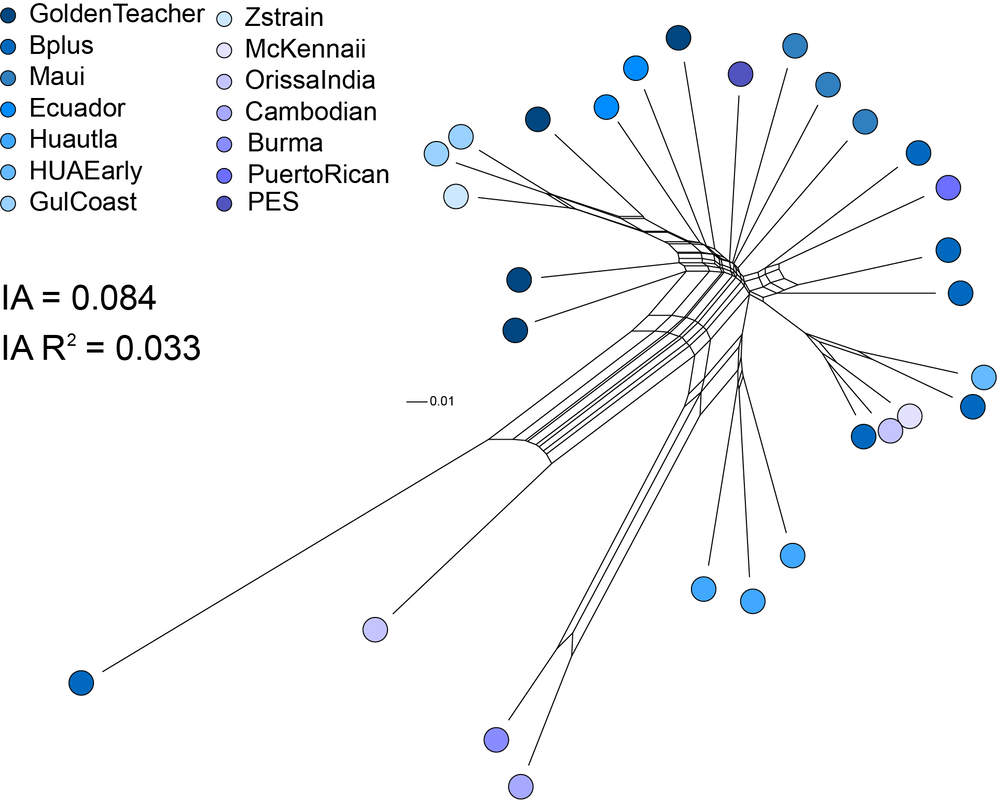
Recommend watching the talk on Clandestine Dimethyltryptamine (DMT) laboratories encountered in Queensland by Dr Tim Currie. The team at Psilopedia have released genomes of 84 cultivated lines of Psilocybe cubensis. How exciting. I did what anyone with a couple of spare genomes of P. cubensis and access to high performance computing would do to determine how the Oz population was related to what people are growing overseas. The Psilopedia genomes are based on DNA extracted from spore prints, which means there are thousands of haplotypes in the assembly. Not great for contiguity but still fantastic for analyses of genetic diversity. I annotated 80 of the Psilopedia genomes and three of our Australian haploid genomes (collected from the Gold Coast). I called SNPs from protein coding regions of sites that were present in every assembly. The SplitsTree figure above is from over 500,000 SNPs and it shows that the Gold Coast population of P. cubensis is completely different to the popular cultivars used by the community. Another interpretation is that there is more genetic diversity in the three haploid genomes from Australia than the entire population of cultivated Golden Teacher (based on branch lengths of genetic distance). Likely, gold tops were introduced to Australia before P. cubensis was widely cultivated. I further looked at the genetic diversity in the Golden Teacher population. This is open to interpretation and would be better with other comparative populations. I interpret this relationship as inbreeding, with little reticulation toward the tip ends and closely related individuals on longer branches. The Index of Association here is a little meaningless at the moment. In obligate outcrossers we expect lower linkage among sites. The Index of Association gets smaller when there is sexual reproduction (increased randomness of allele combinations across the dataset) and higher in clonal populations (the same alleles are always recovered together).
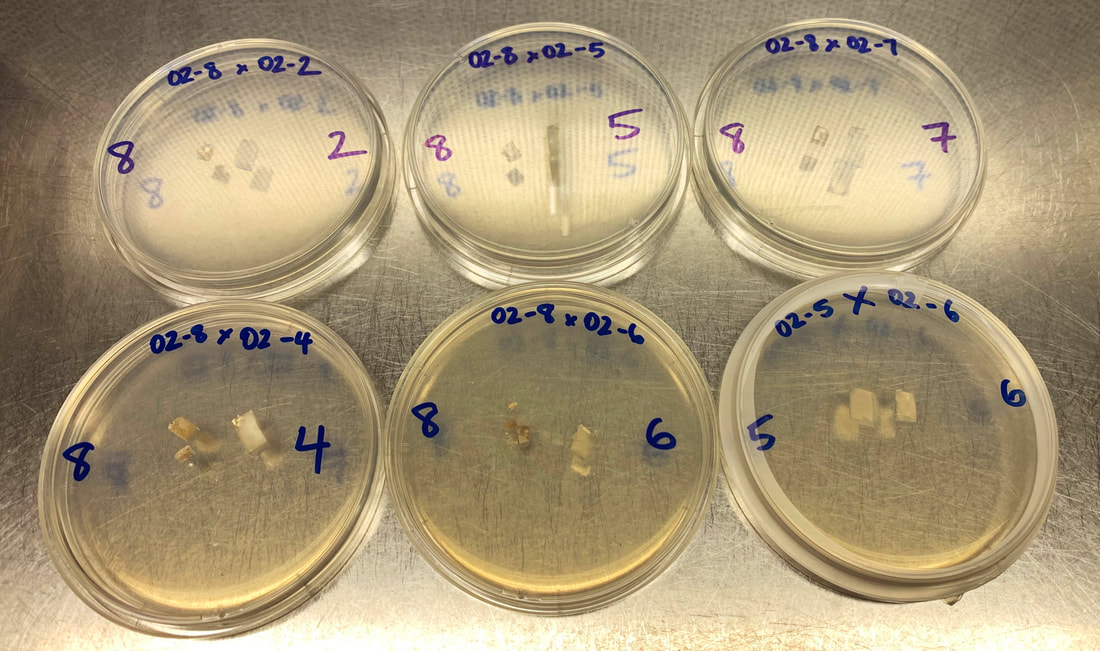
Stay tuned. Next week I will submit another 32 isolates of P. cubensis to sequence genomes from NSW and Queensland and determine whether the population is from one introduction, whether there is outcrossing, the age of introduction and whether there have been any escapes from people cultivating mushrooms. This post contains two updates, (i) a part on the hallucinogenic species in the Northern Territory, and (ii) how to set the mood for fungal reproduction. There was some media attention after the last blog post. It was refreshing to see how many people are interested in fungi, not just magic mushrooms. Thank you to all who have volunteered time to help the project, this means a lot. I looked for names to apply to the species in Kakadu and Litchfield National Parks in case something was a clear match. No name can be applied with 100% confidence, however, when Gaston Guzman described Psilocybe brunneocystidiata from PNG, he hypothesised it would be a close relative of P. yungensis, which was described as a wood decomposer from Bolivia. Is it a coincidence that the mushroom from the NT was sister to P. yungensis in the exact relationship the most prolific Psilocybe-taxonomist predicted for P. brunneocystidiata? We may never know for certain whether this name could be accurately applied to the taxon in Kakadu, but it is a starting point considering we don't have the fungus in hand. I have received emails aplenty of magic mushrooms occurring in the NT, including in Kakadu. Some of these we might suspect are Panaeolus cyanescens (based on their described appearance and their niche in buffalo dung). If anyone from the Mirarr people know about traditional use of mushrooms and they have knowledge that is appropriate to share, we should get in touch. One last point, a new species in taxonomy is one that does not have a binomial Latin name. I accept hypotheses of a clock-like rate of speciation (in which species are approximately 500,000 – 5 million years old), technically there are no 'new' species unless humans have caused selection pressure or new pathways for outbreeding/hybridisation. To say something is a 'new species' is taxonomic jargon for 'let's give this thing a name so we can improve communication'. If you have watched some of my recent seminars (aren't you lucky), you'll have your finger on the pulse with the mating (MAT) genes annotated from the assemblies of P. subaeruginosa. By chance, the population I sampled has a perfect distribution of MAT alleles so that I can confirm our predictions about mating from the genomes. Basidiomycota (mushrooms, smuts, and rusts) have homeodomain genes (usually two of them close together on one chromosome) and pheromone/receptor genes used for mate recognition and signalling. These genes will become useful when we study whether P. cubensis and P. subaeruginosa are native to Australia because we expect a lot of diversity at homeodomain genes in natural populations. Generally opposites attract for sexual reproduction and two haploid fungi must differ at their MAT loci for a compatible cross. If they must differ at both HD and P/R loci, mating is controlled by two loci and different MAT alleles are needed (tetrapolar mating). If fungi only need to differ at their HD locus for a compatible cross, or the HD and P/R loci are closely linked (usually by proximity), only one locus controls compatibility (bipolar mating). Tetrapolar mating is useful for fungi that cannot move long distances because it reduces the chances of inbreeding. Psilocybe subaeruginosa has three P/R loci, each of which segregated in our population. Now we need to work out which of these is controlling compatibility. Here are my predictions for the crosses in the image below based on what we know from the genome. Successful mating will be 8x5 and 8x6, the rest probably will be incompatible. |
Designer Shrooms @ Funky Fungus on 1st July 2023
I started a gig at Funky Fungus as Chief Scientific Officer to make designer shrooms Our research on Psilocybe
|