|
0 Comments
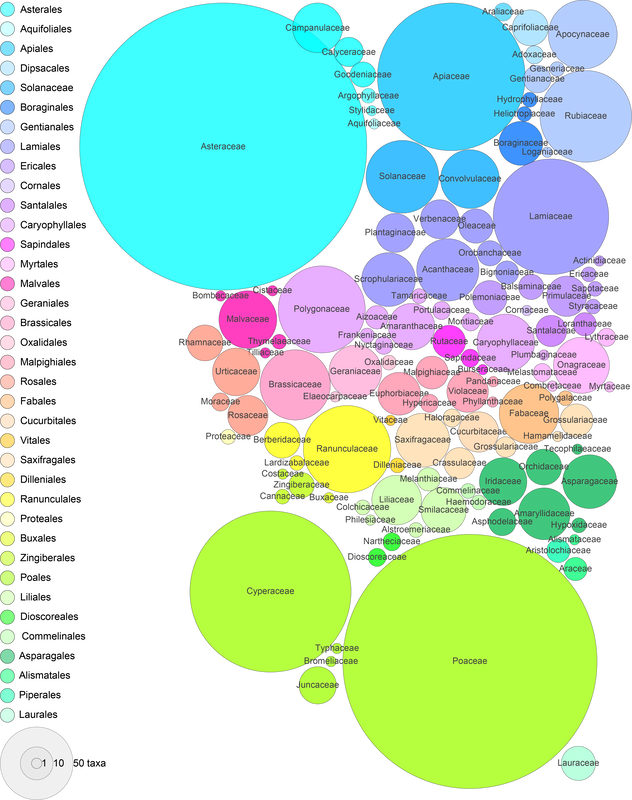
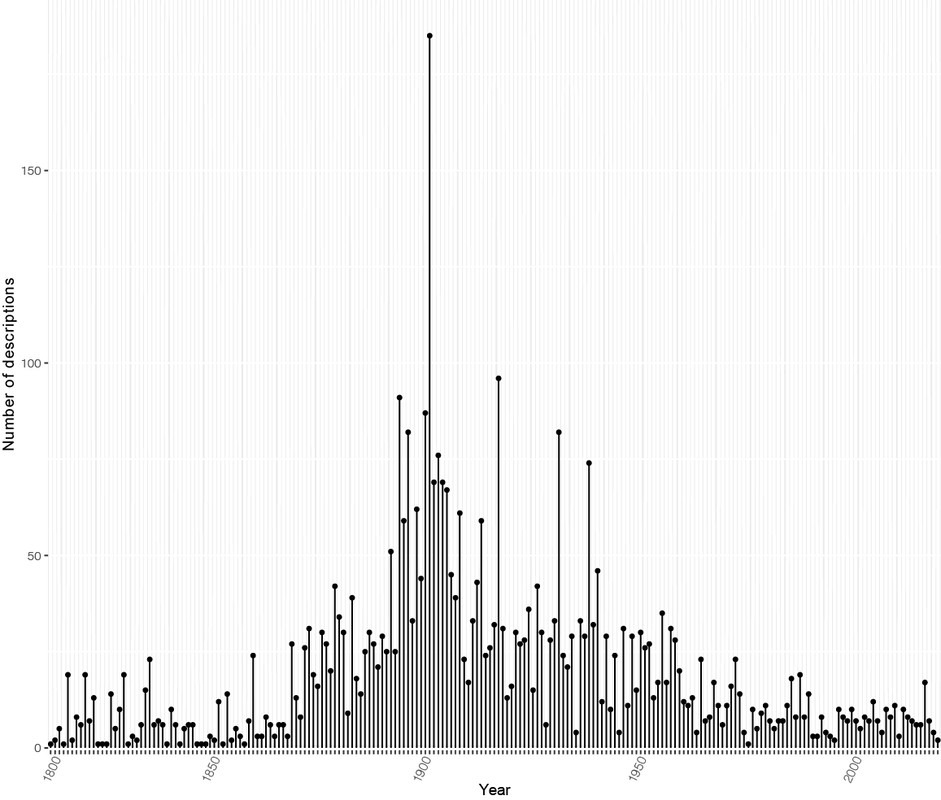
Here's a breakdown of all names described for rust fungi. Just over 10,000 names used. Some will be synonyms, some await description... but the rate of description has certainly slowed (for Puccinia at least).
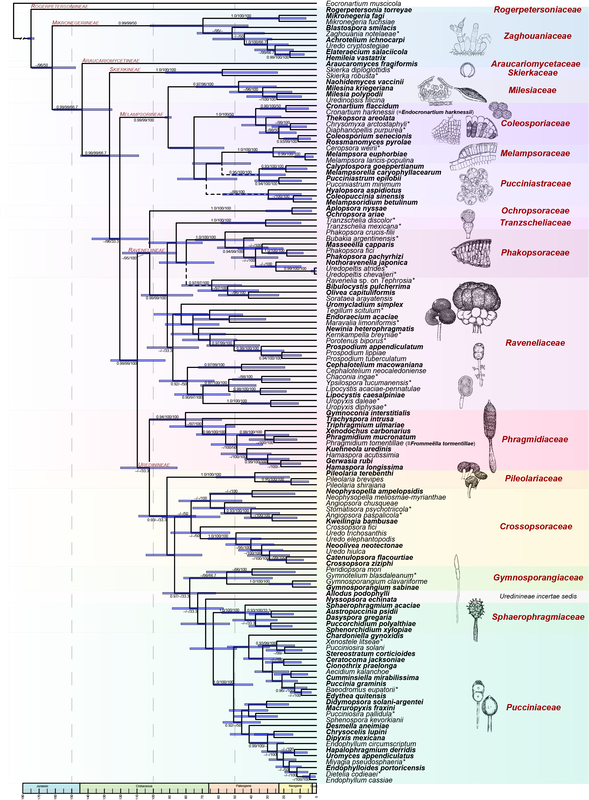
This tree took 10 years. Alignments available on TreeBase and Research Gate. This is a good starting point for anything unknown in rust taxonomy.
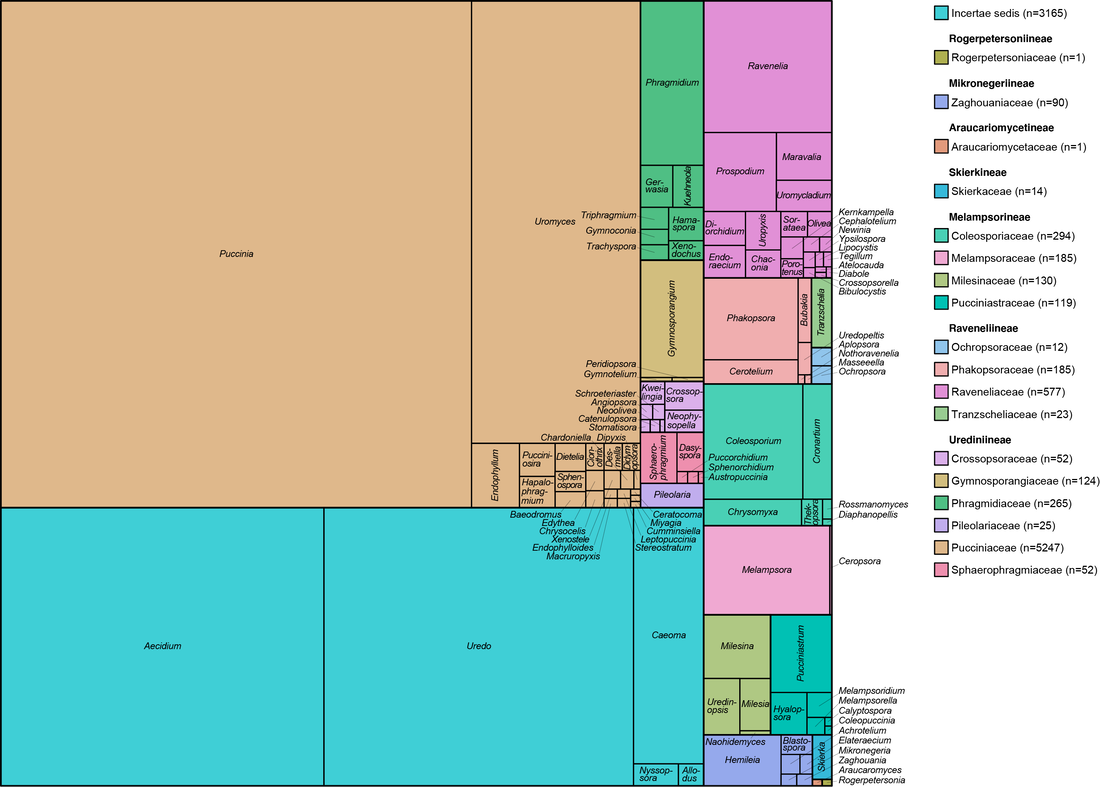
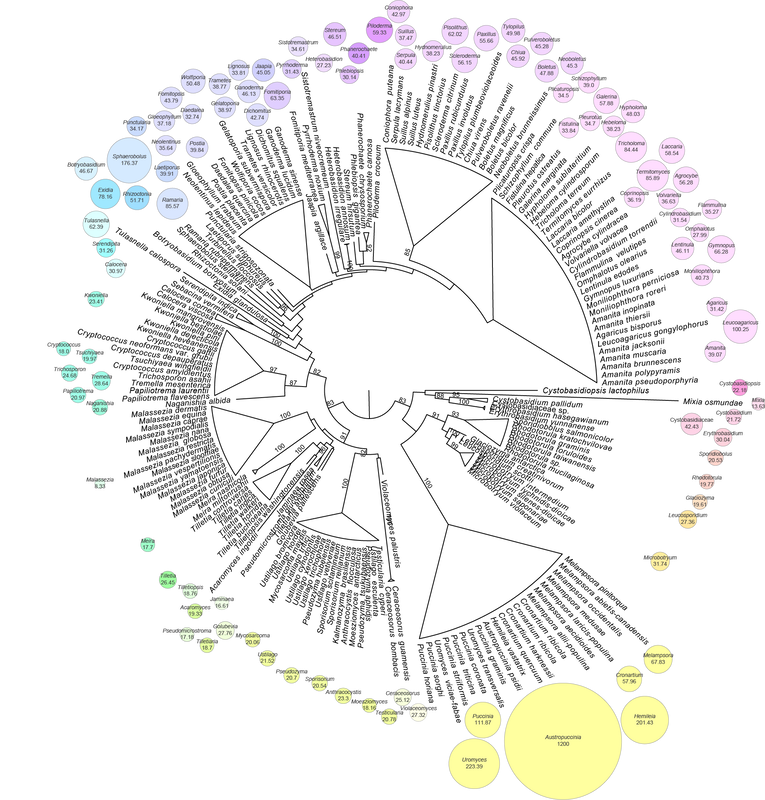
It's pretty cool that rust taxonomists have agreed about familial (or the equivalent) placement of most taxa. These figures illustrate the genera that nobody really agreed about, and with good reason, they're dang tricky to place based on morphology. We know that Hemileia (being a group of tropical, rainforest, autoecious, suprastomatal rusts) is in Zaghouaniaceae, a family sister to other rusts. Sphaerophragmium and Phakopsora both wound up in their own families. Gerwasia on Rosaceae is a classic case for Phragmidiaceae, and Desmella (a taxon on a fern) belongs to the Pucciniaceae. Knowledge on Skierka coming soon and these figures now accepted in FuSE!. (Figures were made in R with ggalluvial).
We sequenced and assembled the genome of the South African strain of myrtle rust and found it was 1.2 Gb (McTaggart et al. 2018). The largest fungal genome assembled so far. Since then I've sequenced the genomes of a few other rust fungi using 10X with Chromium libraries. There'll always be some contamination from host and adjacent bacteria that could skew my early results, but I'll share them anyway, just be mindful things could change.
Puccinia paullula, a suprastomatal rust on Monstera deliciosa, had an estimated genome size over 3 Gb (no doubt a repetitive genome with an N50 of 27 Kb for the contigs). Hamaspora acutissima had an estimated genome size over 6 Gb, with a very high repetitivity index (78.89 %). Skierka agallocha, which has the most chance of contamination because the pustules are embedded in the host, had an estimated genome size of 1.97 Gb. This one is a very fragmented assembly, probably because of the nature of the material we were working with; getting long molecules of DNA in the extraction was a challenge. Aecidium fragiforme, one of my all-time favourite rusts, and one of the most ancestral, had an estimated genome size over 5 Gb. Coleosporium plumeriae was tiny in comparison to all these, only 381 Mb and a low repetitivity index (13 %). Sphaerophragmium acaciae, the sister taxon to myrtle rust, was first estimated at 200 Mb, but I'd used a small amount of reads. When I used the full dataset the genome was estimated over 2 Gb... so a little bit of work to do yet. All these genomes are courtesy of the ABRS and my friend Louise Shuey, who did all of the extractions. If you're interested in any of these data let me know and I'll cut you a piece, there's plenty of data to go around, and the taxon selection is pretty killer (just saying).. My opinion on Mycosarcoma and Ustilago Mycosarcoma tubiforme Mycosarcoma tubiforme Ustilago was named for the burned appearance of an inflorescence infected by smut fungi. The genus was typified by U. hordei (Clinton 1906), and over the last century it has typified taxonomic ranks up to the Ustilaginomycotina. Ustilago became a catch-all for many species of smut fungi on grasses (McTaggart et al. 2012b), and although it is a polyphyletic genus, Ustilago s. str. includes smut fungi that destroy the inflorescence and leaves of Pooid and Chloridoid grasses (Stoll et al. 2005; McTaggart et al. 2012c). Mycosarcoma maydis (corn smut) was long-treated as a species of Ustilago. Brefeld (1812) described Mycosarcoma because M. maydis formed hypertrophied, localized sori and was not congeneric with Ustilago. The relationship between corn smut and other species of smut fungi has been well established in systematic studies (Piepenbring et al. 2002; Stoll et al. 2005; Vánky and Lutz 2011; McTaggart et al. 2012a). McTaggart et al. (2016) used Mycosarcoma for a monophyletic group of smut fungi on grasses with localized, hypertrophied, host-derived sori. This taxonomic hypothesis highlighted that M. maydis was not closely related to other species of Ustilago, and we should expect differences in biology, reproduction and host range between taxa based on their generic classification. The scientific community that work on smut fungi are still able to use Ustilago maydis under the taxonomic proposal of McTaggart et al. (2016). After all, taxonomy is about communication and well-known names should be long-lived. The main consequence/benefit of using Mycosarcoma is that corn smut is distinguished from other species of Ustilago at a generic rank. Thines (2016) proposed that the type of Ustilago should be changed from U. hordei to M. maydis. The proposal was based on a taxonomic and a nomenclatural reason: U. hordei was not the type of Ustilago, and M. maydis was a more important taxon. Clinton (1906) used a footnote to typify Ustilago with U. hordei, and there is no taxonomic basis for the proposal, as outlined by McTaggart et al. (2016). Thines (2016) hypothesized that a taxon was more important than another if it had more hits in a Google search. This was the reason to make a nomenclatural change to the type species of Ustilago. Mycosarcoma maydis represented 75 % of the hits for Ustilago in a Google search (Thines 2016). A nomenclatural decision based on hits in a Google search sets an interesting precedent. Types could be changed on their convenience and popularity as model organisms. For example, Tilletia indica (237,000 hits) is more important than the type species T. caries (46,000 hits); Melampsora lini (136,000 hits) is more important than the type species M. euphorbiae (19,900 hits); Pseudocercospora fijiensis (25,000 hits) is more important than the type species Ps. vitis (21,000 hits); Peronospora destructor (48,800 hits) is more important than the type species Pe. rumicis (2,950 hits), yet the type should have been conserved with the popular Pe. parasitica (82,800 hits) rather than description of this taxon in Hyaloperonospora. Ustilago hordei, reflects the original type species of Ustilago sanctioned as the alpha variety by Persoon (1801), lectotypified by Clinton (1906) and used for over a century by the mycological community (Vánky 2011). Should nomenclature change on a whim to reflect how often the scientific community uses a taxon (at one point in time U. hordei was more important than M. maydis)? If yes, where is the limit for nomenclatural changes based on this reasoning? I think nomenclatural changes should be made in well-reasoned cases with many lines of argument, rather than whichever taxon is in vogue at the time. *Google searches were performed on the 22nd March 2018. Brefeld O (1812) Untersuchungen aus dem Gesammtgebiete der Mykologie. Vol. 15., vol 5. Die Brandpilze und die Brandkrankheiten. Commissions-Verlag v. H. Schöningh, Münster Clinton GP (1906) Order Ustilaginales. North American Flora 7:1–82 McTaggart AR, Shivas RG, Boekhout T, Oberwinkler F, Vánky K, Pennycook SR, Begerow D (2016) Mycosarcoma(Ustilaginaceae), a resurrected generic name for corn smut (Ustilago maydis) and its close relatives with hypertrophied, tubular sori. IMA Fungus 7 (2):309–315. doi:10.5598/imafungus.2016.07.02.10 McTaggart AR, Shivas RG, Geering AD, Callaghan B, Vánky K, Scharaschkin T (2012a) Soral synapomorphies are significant for the systematics of the Ustilago-Sporisorium-Macalpinomyces complex (Ustilaginaceae). Persoonia 29:63–77. doi:10.3767/003158512x660562 McTaggart AR, Shivas RG, Geering AD, Vánky K, Scharaschkin T (2012b) A review of the Ustilago-Sporisorium-Macalpinomyces complex. Persoonia 29:55–62. doi:10.3767/003158512x660283 McTaggart AR, Shivas RG, Geering AD, Vánky K, Scharaschkin T (2012c) Taxonomic revision of Ustilago, Sporisoriumand Macalpinomyces. Persoonia 29:116–132. doi:10.3767/003158512x661462 Persoon CH (1801) Synopsis Methodica Fungorum, vol 1. Henricus Dieterich, Germany, Göttingen Piepenbring M, Stoll M, Oberwinkler F (2002) The generic position of Ustilago maydis, Ustilago scitaminea, and Ustilago esculenta (Ustilaginales). Mycological Progress 1 (1):71-80. doi:10.1007/s11557-006-0006-y Stoll M, Begerow D, Oberwinkler F (2005) Molecular phylogeny of Ustilago, Sporisorium, and related taxa based on combined analyses of rDNA sequences. Mycol Res 109 (3):342–356. doi:https://doi.org/10.1017/S0953756204002229 Thines M (2016) Proposal to conserve the name Ustilago (Basidiomycota) with a conserved type. Taxon 65:1170–1171 Vánky K (2011) Smut fungi of the world. APS Press, St. Paul, Minnesota, USA Vánky K, Lutz M (2011) Tubisorus, a new genus of smut fungi (Ustilaginomycetes) for Sporisorium pachycarpum. Mycologia Balcanica 8:129–135 |
Archives
June 2022
Categories |