|
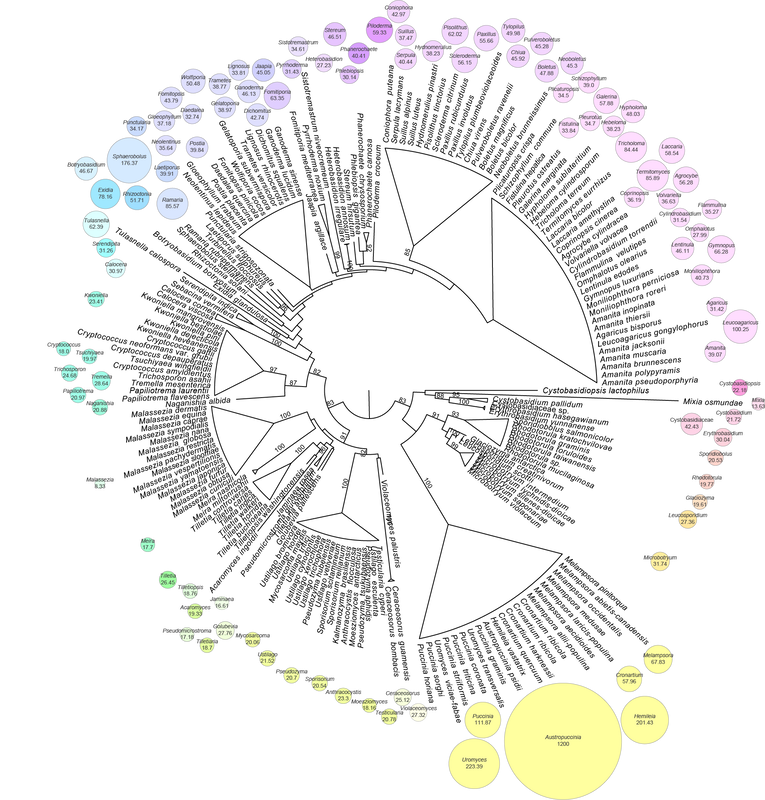
We sequenced and assembled the genome of the South African strain of myrtle rust and found it was 1.2 Gb (McTaggart et al. 2018). The largest fungal genome assembled so far. Since then I've sequenced the genomes of a few other rust fungi using 10X with Chromium libraries. There'll always be some contamination from host and adjacent bacteria that could skew my early results, but I'll share them anyway, just be mindful things could change.
Puccinia paullula, a suprastomatal rust on Monstera deliciosa, had an estimated genome size over 3 Gb (no doubt a repetitive genome with an N50 of 27 Kb for the contigs). Hamaspora acutissima had an estimated genome size over 6 Gb, with a very high repetitivity index (78.89 %). Skierka agallocha, which has the most chance of contamination because the pustules are embedded in the host, had an estimated genome size of 1.97 Gb. This one is a very fragmented assembly, probably because of the nature of the material we were working with; getting long molecules of DNA in the extraction was a challenge. Aecidium fragiforme, one of my all-time favourite rusts, and one of the most ancestral, had an estimated genome size over 5 Gb. Coleosporium plumeriae was tiny in comparison to all these, only 381 Mb and a low repetitivity index (13 %). Sphaerophragmium acaciae, the sister taxon to myrtle rust, was first estimated at 200 Mb, but I'd used a small amount of reads. When I used the full dataset the genome was estimated over 2 Gb... so a little bit of work to do yet. All these genomes are courtesy of the ABRS and my friend Louise Shuey, who did all of the extractions. If you're interested in any of these data let me know and I'll cut you a piece, there's plenty of data to go around, and the taxon selection is pretty killer (just saying)..
0 Comments
Leave a Reply. |
Archives
June 2022
Categories |